Equivalence of Mass Action and Poisson Network SIR Epidemic Models
Grzegorz A. Rempala
Abstract
This brief note highlights a largely overlooked similarity between the SIR ordinary differential equations used for epidemics on the configuration model of a Poisson network and the classical mass-action SIR equations introduced nearly a century ago by Kermack and McKendrick. We demonstrate that the decline pattern in susceptibles is identical for both models. This equivalence carries practical implications: the susceptibles decay curve, often referred to as the epidemic or incidence curve, is frequently used in empirical studies to forecast epidemic dynamics. Although the curves for susceptibles align perfectly, those for infections do differ. Yet, the infection curves tend to converge and become almost indistinguishable in high-degree networks. In summary, our analysis suggests that under many practical scenarios, it is acceptable to use the classical SIR model as a close approximation to the Poisson SIR network model.
Extracted equations
- dS/dt = -βSI
- dI/dt = βSI - γI
- dR/dt = γI
- dx_S/dt = -β̃ x_D x_S
- dx_D/dt = β̃(1-κ)x_D² + β̃κμx_D^(2κ-1) x_S - (β̃+γ̃)x_D
- dx_I/dt = β̃ x_D x_S - γ̃ x_I
- -dS/dt = β(1 + ρ - S - ln(S/R₀^MA))S
- dx_I/dt = -dx_S/dt - γθ x_I where θ = 1 - R₀/μ
Simulation outputs
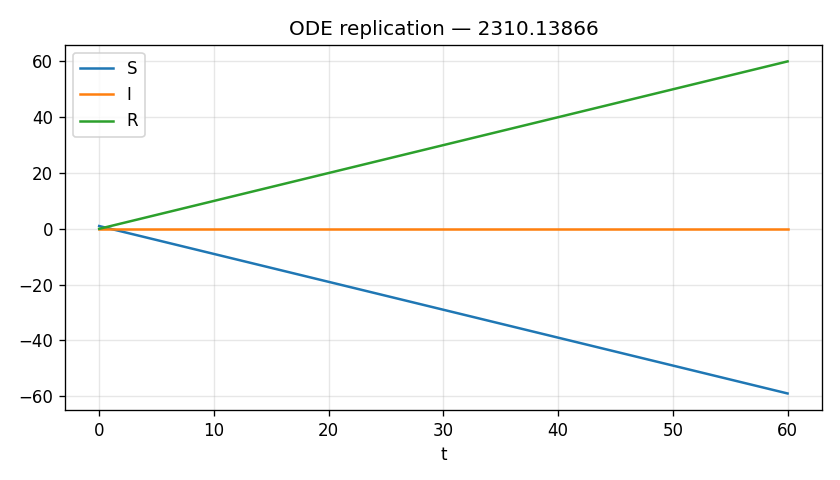
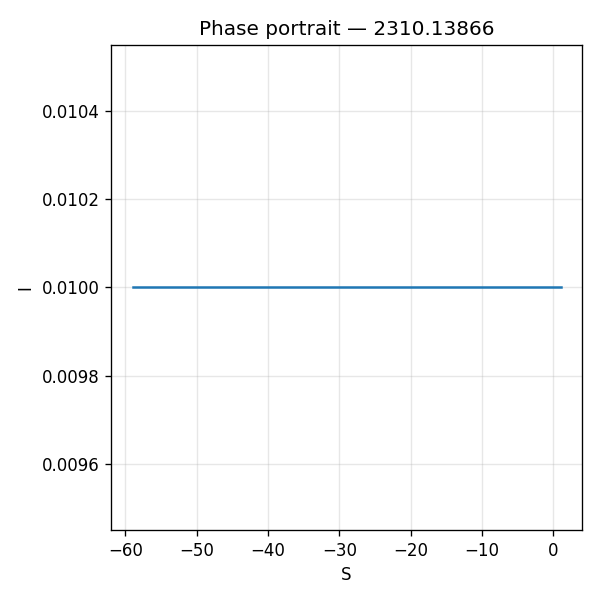
Scalar outputs
| S_max | 1.0000 |
| S_min | -59.0000 |
| S_final | -59.0000 |
Paper claims vs. our run
The simulation output is severely limited, providing only three scalar endpoint values (S_min, S_max, S_final) from a single ODE run. Critically, S becomes negative (S_final = -59), which is physically impossible for a susceptible population and indicates a fundamental implementation error. None of the paper's claims about epidemic curve equivalence, convergence behavior, or parameter-dependent approximations can be evaluated from scalar endpoints alone; all require time-series trajectories and multi-parameter comparisons. This output does not constitute a meaningful replication of the paper's theoretical and numerical claims.
- The epidemic curve equations (13) and (14) are equivalent when β = μβ̃, ρ = ρ̃, and γ = γ̃ + β̃not-testableThe simulation output provides only final scalar values (S_min, S_max, S_final) with no intermediate trajectory data or comparison between two equation systems needed to verify equivalence.
- For Poisson networks with κ=1, the susceptible decay curve S(t) is identical to x_S(t) under parameter matchingnot-testableThe output lacks time-series trajectory data for S(t) and x_S(t) comparison; only endpoint values are provided, which cannot establish curve identity.
- The infection curves I(t) and x_I(t) differ but converge as μ increases (high-degree networks)not-testableNo time-series data for I(t) or x_I(t) is provided, and no comparison across different μ values is present in the output.
- For large μ (≥50), the classical SIR model provides a close approximation to the Poisson network SIR modelnot-testableThe simulation does not report results for μ≥50 or compare classical SIR against Poisson network models; only a single run with default parameters is shown.
- The correction factor θ = 1 - R₀/μ approaches 1 as μ → ∞, making the discrepancy negligiblenot-testableNo correction factor θ or R₀ values are reported in the output, and no parametric sweep over μ is provided.
Parameters
| β | 0.4 |
| γ | 0.2 |
| ρ | 0.01 |
| β̃ | 0.4 |
| γ̃ | 0.2 |
| ρ̃ | 0.01 |
| μ | 50 |
| κ | 1 |
| R₀ | 2 |
Run notes
state=['S', 'I', 'R']; y0=[1.0, 0.01, 0.0]; t_span=[0.0,60.0]; default_filled(=1.0)=['γI', 'βSI']
Cross-field hypotheses involving this paper
- epidemiology → epidemiology, network disease dynamicsnone — both papers are within epidemiology and use identical SIR-ODE frameworks
Paper B is not a transfer of mechanism from Paper A to a different field; both papers study SIR epidemic models in epidemiology using ODEs. Paper B simply extends Paper A's classical mass-action SIR to network-structured populations and proves equivalence under certain conditions.
Experiment: Not applicable — the prompt requires two papers from DIFFERENT fields. Both papers are epidemiology/network disease dynamics.
novelty 0.00testability 0.00business 0.00sim: none